CHAPTER FOUR
Visualizing Prions
Graphic Representations and the Biography of Prions
Jérôme Segal
Institut Universitaire de Formation des Maîtres,
& Centre Cavaillès of the Ecole
Normale Supérieure de
Paris
Eric Francoeur
Max Planck Institute for
the History of Science
“A scientific concept that is not
supported by direct visualization is always difficult to establish, whatever
its origin may be.” D. Dormont
Prions are proteins generally characterized by the ability to exist in
two different forms or more precisely two different three-dimensional
structures, one of them possibly causing disease when it aggregates. The prion
hypothesis, as formulated by Stanley Prusiner, states that this aggregation
causes specific neurological diseases such as Bovine Spongiform Encephalopathy
(BSE). Even if both the mechanisms of this change of conformation and that of
the aggregation are still enigmatic, the prion hypothesis has become a dominant
model to which much heuristic power has been attributed in the 1990s. This
could be a first paradox.
Moreover, whereas three-dimensional structures clearly appear to be at
the heart of the matter, Prusiner used mostly biochemical evidence to develop
his hypothesis, without using, in the early days, any other graphic
representations than that given by electron microscopy. This constitutes the
second paradox at the origin of the present paper since only computer
representations of three-dimensional structures can explain and justify the
prion theory as a model. Here, models are defined as theories with two distinct
properties. First, models have an explanatory power more or less confirmed by
experimental evidence, which distinguishes them from mere hypotheses. Secondly,
models can be applied in domains other than those where they come from. Such
application is possible due to the underlying formalism of models, or, as in
the prion case, to the diffusion of a specific visualization culture.
Since the second half of the 1990s, many scientific journals have
published three-dimensional representations of prion structures, always in the
non-pathological form (the structure of the other form remaining as yet
unknown). What are these representations supposed to bring? Why were they
published, sometimes on the cover of prestigious journals? How have they been
obtained? The use of computers has of course been decisive but more generally,
did the visualization aids provided by bioinformatics help to change the
epistemological status of the prion hypothesis?
To tackle these questions, we will first outline the context in which
three-dimensional structures of proteins, historically called ‘tertiary
structures’, have become an important scientific topic. We will then review the
place of graphic representations in Prusiner’s work, and show how the prion
hypothesis has changed metaphors in biology. This will allow us to concentrate
on the conformational change, and to end our narrative with an analysis of the
main on-going projects on the tertiary structure, with particular emphasis on
the case of ‘yeast prions’.
I
– An overview of molecular
visualization
Until fairly recently, historians and
philosophers of science paid scant attention to the issue of visual
representation in science.[1] Since
the mid-1980s, scholars in science studies have become increasingly concerned
with the role of visualization and visual representation in the development and
practice of science.[2]
From this literature has come the clear conclusion that visual representation
is far from an epiphenomenon of scientific practice, but rather one of its
intrinsic elements.
The issue of visual representation can be
understood not only in terms of techniques and technology but also in terms of
the various practices and activities associated with making ‘natural’ objects
observable and intelligible. Ethnographic studies of laboratory activities have
been particularly instructive in that regard, showing the transformation over
time of research objects and their gradual shaping into pictorial data and graphic
displays.[3] The
present paper deals with visualization in molecular biology.
The recent completion of the Human Genome Project in 2001 has brought
disappointment for all those who believed it would lead to rapid progress in
gene therapy or at least provide a better understanding of protein synthesis.
This worldwide project has produced the complete sequence of human DNA. As is
well known, this DNA sequence codes for the amino acids that constitute
proteins. The ‘primary structure’ of proteins is given by this sequence of
amino acids, and the assembly of some regular structure, such as alpha helices
and beta sheets, defines the ‘secondary structure’.[4] If these structures can be
defined without complex graphic representations, the full ‘tertiary structure’
of proteins, their functional three-dimensional shape, cannot be easily
described in its full complexity without visualization devices since the
secondary structure only gives hints to the arrangement of the tertiary
structure (only parts that are identified as helices etc.). The problem of
‘protein folding’ corresponds to the process during which the protein acquires
its tertiary structure.
I-1. The emphasis laid on DNA in the ‘central dogma’
In 1957, four years after publishing with James Watson the structure of
DNA (1953), Francis Crick held a conference “on protein synthesis” (Crick
1958).[5] He clearly stated why he
chose to concentrate on the primary structure:
Our basic handicap at the moment is that we have no easy and precise
technique with which to study how proteins are folded, whereas we can at least
make some experimental approach to amino acid sequences. For this reason, if
for no other, I shall ignore folding in what follows and concentrate on the determination
of sequences. (Crick, 1958: 144)
Even if he insisted at the beginning of his talk on the fact that, as in
the case of enzymes, proteins owe their specificity and activity to the
properties of their tertiary structure, Crick dealt mostly with ‘information’,
which he defined as “determination of sequence, either of bases in the nucleic
acid or of amino acid residues in the protein”. He developed his views under
the hypothesis that “folding is simply a function of the order of the amino
acids” and the conference became famous because of the formulation of what he
called “the Central dogma” of molecular biology. As he wrote, “This states that
once ‘information’ has passed into proteins it cannot get out again”, which
means that information cannot flow from proteins to genes (Crick is somewhat vague
about the role of RNA which determines the sequence of amino acids).
Crick later explained that he meant ‘axiom’ rather than ‘dogma’ but the
diffusion of this idea led many researchers to consider the analysis of DNA as
a quest for the Holy Grail. Under the assumption that DNA sequences would
explain protein synthesis, most funding went to genetics and protein studies
became somewhat neglected. A static conception of protein dominated, whereas
biochemists knew that the study of the folding process requires a dynamic
approach.
I.2. Interactive molecular graphics and the heuristic role of computers
Molecular biology has been developed in a civilization based on writing.
Scientists publish articles and even when they talk, they say that they present
‘papers’. How did they acquire and transmit their knowledge regarding protein
tertiary structures? To understand the structure of molecules, biologists
managed – often with the aid of X-ray analysis – to build physical models of
the molecule they wanted to study. Robert
Corey and Linus Pauling, who offered tools to identify secondary structures,
designed various types of models. Some of these models emphasised the volumes
occupied by atoms in the molecules and afforded an understanding of steric
hindrance. The Corey-Pauling-Koltun (CPK) space-filling models, based on an
original design by Corey and Pauling, became very popular in the late 1960s
(Francoeur 2001).
These physical models did not allow for
satisfactory manipulation and their construction often proved physically
impossible for big molecules (some biologists even contemplated building models
under water to avoid the effect of gravity). The breakthrough came with the
development of time-shared mainframe computers, which allowed real-time
functioning and interactivity between the user and the machine. The precise origin of the concept
and techniques of interactive molecular graphics can be traced back to a group
of scientists around the molecular biologist Cyrus Levinthal
(1922-1990), active at the MIT in the mid-1960s (Francoeur
& Segal 2004).
Interactivity was the key element of his visualization device called the
‘Kluge’. It referred to the relative ease with which the scientist was able to
transform the display to highlight particular features of the displayed object
or modify specific parameters of a simulation and get a fast or immediate
visual feedback. In short, this interactivity implied a capacity to experiment
and tinker with the data being modelled or the phenomenon being simulated.
Skilled scientists learned to see what was being disclosed and in this sense,
interactive molecular graphics became a way of revealing the inner character or
hidden nature of things.
Because the Kluge was a vector-based display, only the bonds between the
atoms could be represented, recreating the visual experience of skeletal models, without the problems of gravity. The
illusion of three-dimensionality was created by rotating the structure on the
screen and having the user control the rate of rotation through the
“track-ball”. A light-pen and buttons were also used to interact with the
displayed structure.[6]
Many different programmes and visualization devices were developed
between the 1960s and the early 1980s when Prusiner introduced the prion
hypothesis. In the mid-1970s for instance, a first protein structure was solved
by means of crystallography and visualized entirely with computers (without
building a physical model).[8]
The scope of this paper does not allow us to comment on the place of
visualization in all the different works on protein structure. The important
characteristic of the history of molecular visualization is that a co-evolution
exists between the state of the knowledge and the representation of structures.
For example, when the relevance of describing secondary structures with alpha
helices and beta sheets was admitted, schematic
conventions to represent these structural elements were adopted. In this
sense, we will try to show in the following sections how representation
determines current knowledge related to prions, keeping in mind that specific
modes of visualization “frame” the thinking about the object they represent. Time
has now come to look at how prions have been
represented, bearing in mind that representations are a product of scientific
activity and also influence the way science is being done. Our aim is to see
how these representations affect prions as epistemic things, which at the same
time result from investigations and steer their course, until they finally
settle into well-defined concepts.[9]
II – Representations of Prusiner’s prion hypothesis
II-1.The function of representations in Prusiner’s work in the 1980s
In his 1982 publication in Science,
Stanley Prusiner introduced the word ‘prion’ to denote “small proteinaceous
infectious particles”. The methods he used belonged to a large extent to biochemistry
and also to virology for the study of infectious properties (Prusiner 1982).[10] The major question
concerned the way in which prions “replicate”, if they are devoid of nucleic
acids. An “interesting analogy” was made with retroviruses (where in a schematic
way, information flows from RNA to DNA), and also with the “auto catalytic”
property of the tobacco mosaic virus. For the most part, Prusiner’s theory was
based on the long observation of diseases like scrapie, kuru and
Creutzfeldt-Jakob disease (CJD). Determining the molecular structure of prions
was only considered as a means to gain better understanding of the aetiology of
these diseases: “A knowledge of the molecular structure of prions may help
identify the aetiology of some chronic degenerative diseases of humans.”
(Prusiner 1982: 143). At that time, the idea that a molecule could exist in two
conformations, one of them being able to aggregate, was not mentioned.
In a review article Prusiner published in 1984, the main issue was still
the replication or reproduction of prions in the absence of nucleic acids
(Prusiner 1984).[11]
The question asked in relation to this “biological conundrum” was nothing more
than “what is the nature of their genome?” (Prusiner 1984: 48). Prusiner had
tried to isolate the infectious agent and produced pictures. On the second page
of the paper, we find these two micrographs:
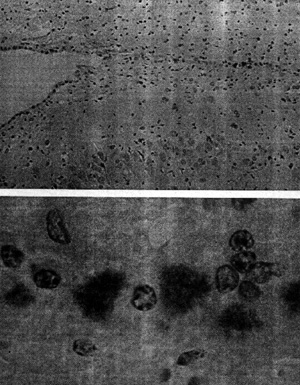
Figure 1 (from Prusiner 1984: 49)
with the following caption:
“Prions in the brain of a
hamster are identified by an immunological staining technique. The hamster had
been infected with scrapie, the prototypical prion disease, which in nature
affects sheep and goats. After an incubation period of roughly two months a
section of brain tissue was exposed to antibodies with a specific affinity for a
protein called PrP, the major constituent of the prion, and possibly the only
constituent.” (emphasis in original)
This text is worth quoting entirely since it raises the question of the
nature of the observable: it is not prions that are directly observed but only
antibodies, which have the specific property of reacting with prions. The
antibodies used in the preparation were labelled with an enzyme (peroxidase)
which, as Prusiner explained, “catalyses the conversion of a colourless reagent
into a dark stain.” (Prusiner 1984: 49). The micrographs showed these stained
structures which were actually not ‘prions’. Electronic microscopy was used to
show what was thought to be “aggregations of prion ‘rods’”, described as “tufts
with a fluffy texture”. These rods were supposed to be “a condensation of
perhaps 1,000 PrP molecules” and the fact that they were indirectly represented
(with specific antibodies) helped to stabilize the theoretical existence of
prions as infectious agents devoid of nucleic acids. Thanks to these
micrographs, the main issue could shift from the search for nucleic acids to
the structural study of these rods, which were noticed to “closely resemble
amyloid plaques”, specific to diseases such as Alzheimer’s. A scientific
culture related to visualization devices then emerged not only in Prusiner’s
prion research but more generally in the TSE field.[12]
As shown by
A step further was then made in the structural analysis of PrP. In a
paper published in 1988, Prusiner indicated that PrP was made of 254 amino
acids, and concluded that “defining the chemical and/or conformational
differences between PrPc and PrPsc is of paramount
importance, as is learning how to synthesize biologically active prions” (Prusiner
1988: 117). At that stage, the existence of prion nucleic acid was qualified as
‘hypothetical’ and Prusiner proposed a diagram to visualize three main
hypotheses of the conformational change: a) the existence of prion nucleic
acid, b) the modification by PrPsc of the gene encoding PrP, or c)
the self triggering of PrPsc to induce a conformational change in
PrPc:
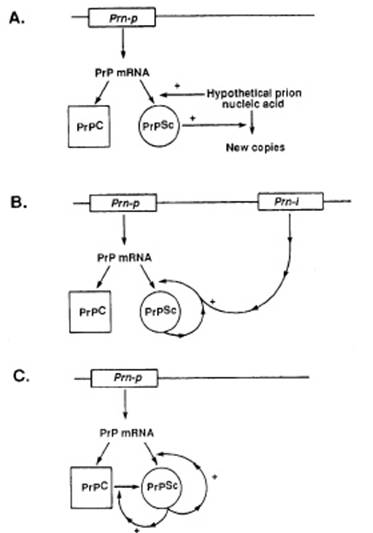
Figure 2 (from Prusiner 1988: 121)
The caption indicated that this diagram illustrates “three possible
models of prion multiplication” (to be compared with the four models of another
1988 paper from the Prusiner group, figure 3 of Poulsen & Andersen paper in this volume, which also
represent genes and enzymes). The word ‘multiplication’ instead of
‘reproduction’ or ‘replication’ previously used, clearly showed that the first
hypothesis was given less and less credit. Poulsen
& Andersen showed how another diagram published in 1991 helped Prusiner illustrating his theory (Figure 1 of their chapter
in this volume).
II-2. Trying to predict the secondary and
tertiary structures to understand the conformational change
The secondary structure of PrP was proposed by Prusiner and his team in
1992, based on biochemical models (Gasset et
al. 1992). Prusiner and colleagues used synthetic peptides reproducing the
four parts of PrP in which they hypothesized the existence of a-helical regions. Three out of the
four synthetic peptides formed amyloid plaques composed largely of b-sheets. Hence, Prusiner and
co-workers came to the idea that the putative conformational change between PrPc
and PrPsc was due to a change of a-helices into b-sheets.
Micrographs were shown to illustrate the authors’ hypothesis on the
secondary structure. The regions that “might form a-helices under monomeric conditions” were
designated H1, H2, H3, and H4. The caption read: “Electron micrographs of H1(…),
H3, H4”). In fact, this caption was rather misleading since only aggregations
of polymerised peptides were displayed:
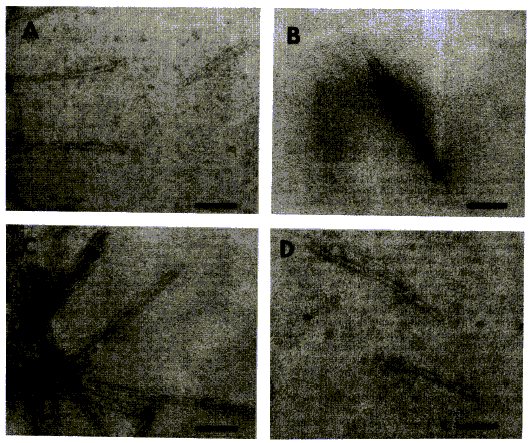
Figure 3 (from Gasset et al. 1992:
10943)
One may wonder about the relevance of reproducing these micrographs.
Were they regarded as visual proof that PrPc contained three or four
helices that could change into b-sheets in PrPsc? Since they did not demonstrate it, one
could argue that they actually weakened this theory.
In any case, the prediction of the secondary structure of PrPc had two important consequences. On the one
hand, it allowed Prusiner and colleagues to better characterize the change of
conformation. In the same year, the diagram used to illustrate the
multiplication of PrPsc was much more univocal compared with the
previous one (see Figure 2):
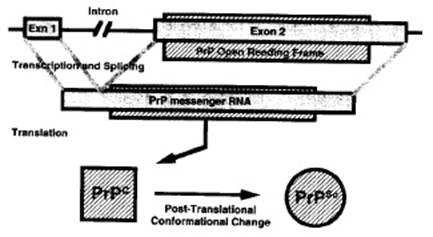
Figure
4 (from Prusiner
1992: 12278)
On the other hand, Prusiner’s prediction of the secondary structure of PrPc shifted the debate away from the notion
that PrPc could replicate without nucleic
acid. Scientists then started to focus on this prediction, and some disputed
it.[13]
In 1992, a specialist in computational biology, Fred E. Cohen, had joined
Prusiner’s team. In 1994, two years after the
publication of the predicted a-helical regions, they proposed a three-dimensional structure of PrPc
(Huang et al. 1994). In this paper,
the PrP used was common to different species. We read that “PrP amino acid
sequences from 1 avian and 11 mammalian sources including chicken, cow, sheep,
rat, mouse, hamster, mink and human were used.” (Huang et al. 1994: 7139).
Prusiner and colleagues’ prediction of the
three-dimensional structure of this PrPc was
presented as a result of computational studies and for the first time ‘computer
modelling’ featured in the keywords. Following up the 1992 paper, the
researchers tried to explain the stable tertiary structure of PrPc.
The transition from the secondary to the tertiary structure was made by
“exploiting recent advances in protein structure prediction algorithms” in
order to obtain a three-dimensional structure of PrPc “based on a
family of homologous amino acid sequences.” (Huang et al. 1994: 7139).
This notion of homology depended on the constitution of databases like
ExPASy (Expert Protein Analysis System) at the Swiss Institute of Bioinformatics whose Internet server had just
been opened (1 August 1993).[14] When the 1994 paper was
published structure predictions were being tested and compared to
crystallographic and Nuclear Magnetic Resonance (NMR) studies.[15] In December 1994 the
first meeting on Critical Assessment of techniques for protein Structure Prediction (CASP) was organized in Asilomar. The idea was to
organize a contest between computer models, differentiating three topics:
‘comparative modelling’, ‘fold recognition or threading’, and ‘ab initio folding’. Fred Cohen,
co-author of the 1994 paper, was in charge of the last topic (Huang et al. 1994).[16]
In the 1994 paper, the four helices identified two years earlier were
arranged in a three-dimensional structure:
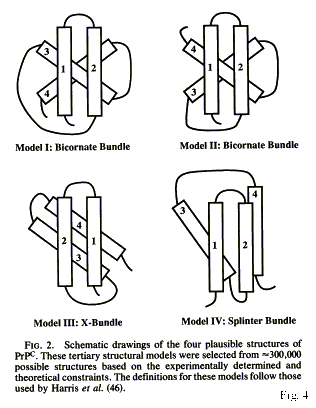
Figure 5 (from Huang et al.
1994: 7141)
Among four possible topological arrangements an X-bundle structure was
chosen based on minimal distances between helices, even though the authors
explained that the algorithms they had used to predict the secondary structure
were probably not appropriate for a protein that exists in two conformational
isoforms.
Based on the X-bundle structure, a three-dimensional structure of PrPc
was proposed by Prusiner and colleagues:
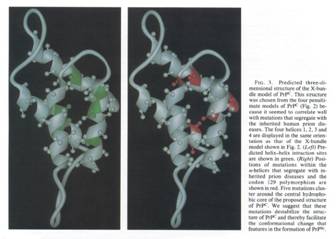
Figure 6 (from Huang et al.
1994: 7142)
In this Figure, we see the same structure twice but with different
indications. On the left-hand side, the predicted helices interaction sites are
highlighted, whereas on the right-hand side, we see the mutation points that
were supposed to explain genetically the difference between PrPc and
PrPsc. This figure was used as a means to understand how the
conformational change could take place. The caption stated: “we suggest that
these mutations destabilize the structure of PrPc and thereby
facilitate the conformational change that features in the formation of PrPsc
(Huang et al. 1994: 7142).
In a review article also published in 1994, the prion was no longer
presented as a hypothesis but rather as a “concept” (Prusiner 1994). The
section “development of the prion concept” was placed just before the section
on the discovery of the prion protein and Prusiner was proud to announce that
“after a decade of severe criticism and serious doubt, the prion concept is now
enjoying considerable acceptance.” (Prusiner 1994: 658).
From the mid-1990s onwards, graphic representations of the
three-dimensional structure played an increasing role in prion research.
Differences in the three-dimensional structures of PrPc and PrPsc
became centre stage. Just as he had proposed a structure of PrPc in
1994, Prusiner proposed a three-dimensional structure
of PrPsc in 1996 (Huang et al.
1996). Prusiner and colleagues first chose amongst six topological
arrangements and then used databases. Two figures in this 1996 paper used the
same conventions as those used two years earlier:
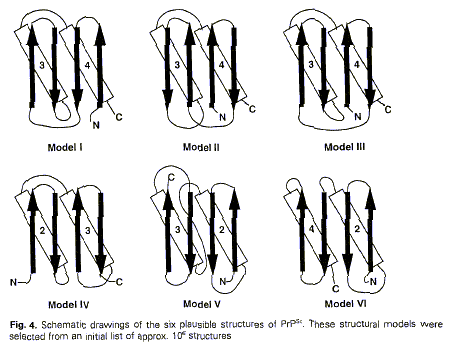
Figure 7 (from Huang et al.
1996: 57-58)
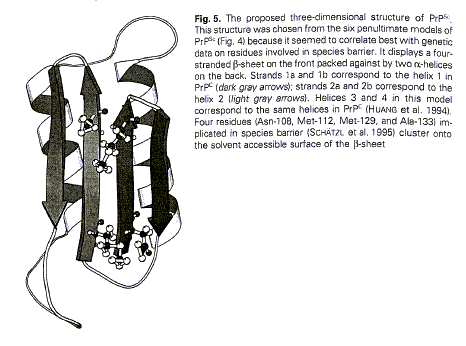
Figure 8 (from Huang et al.
1996: 58)
Visual representations of PrPc and PrPsc led to a better understanding of the
conformational change, linking the results obtained by genetics and
biochemistry to those obtained by computer modelling. The outcome was shown by
superimposing two diagrams, one representing a classical chemical process and
the other representing three-dimensional structures:
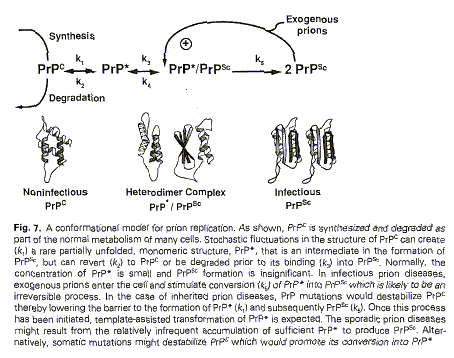
Figure 9 (from Huang et al. 1996: 61)
Figure 9 introduced spatiality to gain a better understanding of prion
structure and compared to Figures 2 and 4 above, marked a transition in the
epistemological function of representations.
To try to explain the conformational change and show more precisely the
possible interactions between parts of the
three-dimensional structure of PrPc, namely helices, Prusiner
decided to reinforce the biocomputational part of his research. In a paper
published in September 1997, he proposed the “Solution structure of a
142-residue recombinant prion protein” of a Syrian Hamster (SHa) (James et al.
1997). Using NMR, Prusiner took advantage of visualization software
developed in
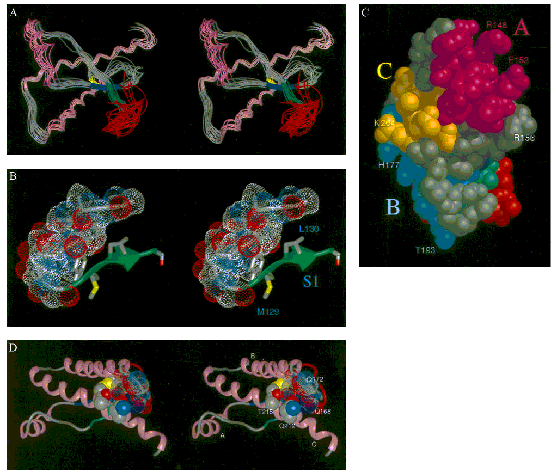
Figure 10a (from James et al. 1997:
10088)

Figure 10b (from James et al. 1997:
10088)
As the original caption of the stereo views indicated, these graphic
representations displayed information that could not be conveyed in textual
format.[19] Such tangling up of alpha
helices and beta sheets was too complex to be described and scientists had to
be trained to decipher stereo views. The analysis of the proposed
three-dimensional structure (Parts ‘D’ and ‘E’ of Figure 10) correlated with a
study of PrPc
in different species led to the formulation of the ‘protein X’ hypothesis,
which would be species-specific and act as a molecular chaperone in PrPSc
formation.[20]
This demonstrates that graphic representations can play an active role in
scientific work since they can lead to new hypotheses and models.
In October 1997, Prusiner was awarded the Nobel
Prize for physiology or medicine. He was awarded alone (which had not happened
for 10 years) and his theory was not yet proven. This gave rise to criticism as shown by Kim (this volume). The fact remains that on the sociological
level the effect of the award was to reinforce the validity of the prion
concept. In his Nobel lecture, Prusiner used the pictures of a modelled
three-dimensional structure displayed above and these images benefited from
widespread exposure (Prusiner 1998).[21]
Soon afterwards, further emphasis was laid on graphic representations. Until
then, protein studies had been dominated by a rather static approach: in line
with Anfinsen’s theories, the folding process was characterized by its initial
and final states (Anfinsen 1973).[22] In contrast, the
existence of a conformational change was now leading to a new dynamical
approach. In terms of visualization, this new approach made the presentation of
results in printed format difficult. Colleagues interested in this kind of work
had to experience on a screen the proposed conformational change. A paper
published in December 1997 reported on the flexibility of a recombinant PrP. In
the abstract, Prusiner and colleagues announced that “detailed information
about PrPc structure may provide essential insights into the
mechanism by which these diseases develop” (Donne et al. 1997: 13452). The announced “detailed information” was
provided in a figure where the flexibility levels corresponded to a colour
scale:
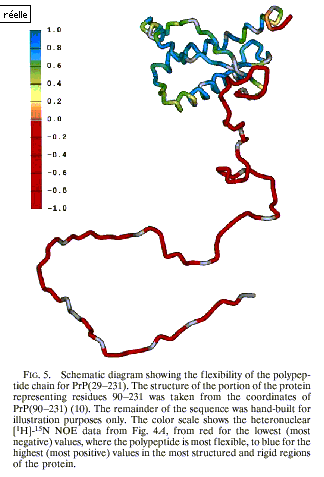
Figure 11 (from Donne et al. 1997: 13456)
The three-dimensional structure of PrPc
was used to convey information aimed at explaining the conformational change in
its dynamical aspects. It is usually
difficult to represent in one’s mind the flexibility of proteins. In the last
five decades, structures have been extensively represented, whereas dynamical
properties were neglected. Though in the early 1960s Levinthal and colleagues
produced films, these were not widely diffused. As a result, when seeing a
protein researchers used to think only in terms of structure. In contrast,
thanks to the use of different colours, Figure 11 represents the ability to
move, that is, readers can imagine the movements of the different parts of PrPc.
In this sense, from the adoption of the dynamic approach, diagrams shaped the
reader’s mind. This exemplifies how representations
of prions have become epistemic things: diagrams are presented as the outcome
of research but they also influence the way researchers define their object.
This brief review has shown that in order to develop and foster the
prion hypothesis, Prusiner increasingly resorted to a range of visual devices,
from micrographs of prion rods to computer modelling of PrP. Colourful graphic
representations generated by computer have featured on the cover of a number of
journals and in this way, the prion hypothesis became so popular that it could
serve as a model. In particular, computer modelling enabled precise
representations that permitted to get a better insight into the conformational
change. In the following section we will see that the progressive diffusion of
this kind of work has changed the way biologists interested in protein studies
and TSEs use metaphors.
III – The use of metaphors
Metaphors have different statuses in science. If some of them are
recognized as such, others derive from the development of predominant
scientific discourses and are used somewhat unconsciously.[23] The development of the
prion hypothesis and the popularisation of related representations provide a
unique opportunity to analyse a shift in the use of metaphors in molecular
biology. The ‘informational metaphor’ has been progressively replaced by other,
more concrete, metaphors like the ‘domino-stone’.
III-1. The power of the informational metaphor
Traditionally the
explanation of the process of infection has been dominated by a discourse based
on information theory. Information theory derived from the mathematical theory
of communication and from cybernetics, and had applications in many different
fields. This development was concomitant with the discovery of the DNA
structure. In the lapse of time between the publication of Crick and Watson’s
paper (1953) on the double helix and the identification of the genetic code
(1961), information theory deeply influenced discourse production in molecular
biology. Kay (2000) has shown that even though on the scientific level
information theory has been of little help, it has nevertheless generated
important informational metaphors. Genetic ‘information’ was at the core of a
number of studies. The discovery of retroviruses in the early seventies (for
which Howard Temin
received the 1975 Nobel Prize) did not really change this conceptualisation
since scientists were still speaking of an information flow, though from RNA to
DNA.
In academic journals
devoted to the history of science, only one article has been published on prion
history, which explores the challenge of the ‘central dogma’ of molecular
biology by the prion hypothesis (Keyes 1999). However interesting it
may be, the discussion is rooted in a misunderstanding of the notion of
information in biology. Keyes seems unaware of the metaphoric nature of
the notion of information. Thus, in addition to the classical “sequential
information” she proposes the concept of “conformational information”: “a
possible new method of replication achieved via the transfer of conformational information forced a
reassessment of the elements of molecular biology’s theoretical framework.”
(Keyes 1999: 4). Unsurprisingly, she also defines the prion as an “information
molecule” and grants it with an “informational role” (Keyes 1999: 210).
In stark contrast, other authors, mostly
scientists, found in the prion theory an opportunity to get rid of this
information metaphor and chose to illustrate their point of view with graphic
metaphors that differ from the traditional arrows illustrating information
flows.
III-2. The ‘domino-stone’ metaphor
As it became clear that thinking in terms of
genetic information was not relevant to the understanding of prion diseases,
other metaphors were developed to explain the spread of PrPsc. In
1996, Adriano Aguzzi at the Institute of Neuropathology (Zurich University
Hospital), and Charles Weissmann at the Institute of Molecular Biology
(University of Zurich), worked on PrPc and showed that this molecule
was required for infection by PrPsc. Studying the “propagation of
the infectious agent”, they gave up the information metaphor and introduced the
‘domino-stone’ metaphor: “Within the framework of the protein-only hypothesis,
these findings [the fact that PrPc is required for the spread of
scrapie] may be accommodated by a ‘domino-stone’ model in which spreading of
scrapie prions in the CNS [central nervous system] occurs per continuitatem through conversion of PrPc
by adjacent PrPsc.” (Brandner
et al. 1996: 13151). This shift was
so important to them that they made a ‘model’ out of it, which has been
extensively used by Aguzzi’s team to study the
conversion of PrPc into PrPsc.[24] In
a paper published in 2000, the domino model illustrated neuroinvasion in the
peripheral neural system. The authors put forward that there could exist “a
mode of transport in which PrPc localized
on the PNS is converted into PrPSc in a ‘domino’ fashion centripetally
towards the CNS.” (Glatzel and
Aguzzi 2000: 2820), and then referred to their 1996 paper.
The progressive acceptance of the prion hypothesis
was accompanied by other metaphors of graphic inspiration.[25] In
textbooks or tutorials the “rotten apple” metaphor is used to illustrate how a
property can spread without information flow. For instance, on a website
devoted to BSE one finds this statement: “Like a rotten apple, once inside the brain, the
mutant form of prion protein turns the native protein into more copies of the
deviant, infectious form”.[26]
Work on the conformational change and the three-dimensional structure of
prions led to a shift from the informational metaphor to these more graphic
metaphors. In turn, the emergence of these new metaphors has contributed to
further stimulate the search for the three-dimensional structure. As a result,
since the mid-1990s knowledge of the three-dimensional structure has become a
holy grail, and not only in Prusiner’s work. As we will see below, many
researchers have now joined this race and different approaches have been
devised.
IV – The race to the tertiary structure of prions: different means for a common goal
Over the years Prusiner’s prion theory gained
widespread acceptance and computer technology soon was at the forefront of
protein studies. Yet, if Prusiner’s work has been
decisive in establishing the prion theory, his papers on prion structure
(mostly on the secondary structure) have not been regarded as seminal. Prusiner became only one contestant among others in the
race to determine the three-dimensional structure of PrP.
In the opening speech of an international conference titled
“postgenomics”, held in July 1998 at the Max Planck Institute for the History
of Science, its director H.-J. Rheinberger stressed that “instead of being
theory-driven, [molecular biology] appears to be eminently technology-driven”.
This statement clearly applies to research on the structure of prions, where
two main techniques have been used, namely, crystallography (see part II) and
NMR.
IV-1.
Crystallography and NMR in prion research
Until the mid-1980s, the main approach to solve three-dimensional
protein structure was through X-ray diffraction analysis, which makes use of
crystallized proteins. Studies by John Kendrew and Max Perutz in the late
1950s, respectively on myoglobin and haemoglobin, were emblematic of the
crystallographic approach. About ten years before, Felix Bloch and Edward
Purcell had come up with the principle of NMR, which allows the detection of
subatomic and structural information of molecules. In NMR a strong magnetic
field (the stronger the field the higher the resolution) is applied to a sample
and measures of how the system responds to radio waves are taken.[27] Initially NMR helped in
chemistry to analyse quantitative mixtures containing known compounds. It took
a long time for it to be applied to biological molecules.
William Dale Phillips (1925-1993) was one of the pioneers in the late
1960s, when the Swiss Kurt Wüthrich arrived at the Bell Laboratories to work on
NMR.[28] Wüthrich initially
focused, as he recalls, “on the metal ion coordination in the active sites of
hemoproteins and on the electronic structure of the heme group” (Wüthrich 2001:
923). It was only from the mid-1970s onwards that Wüthrich tried to apply
NMR to de novo protein structures,
that is, to proteins whose structure is unknown. In the late 1970s, Richard R.
Ernst (1991 Nobel Prize for Chemistry) worked with Wüthrich to develop
two-dimensional NMR techniques. In 1984, NMR proved as useful as X-ray
crystallography to determine structures (Ottiger et al. 1994). In 2001, two
representations of the backbones of the heavy-atoms of a protein the tertiary
structure of which was unknown (a-amylase inhibitor tendamistat), were independently produced with the
two techniques and the graphic representations matched quite well:
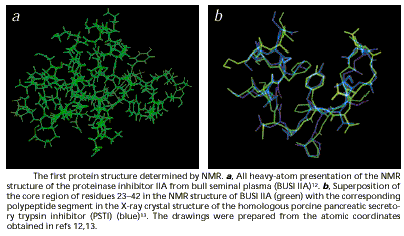
Figure 12 (from Wüthrich 2001: 924)
This was regarded as visual confirmation that
NMR was indeed of interest in molecular biology. In addition, NMR was a kind of
complement to crystallography. Whereas crystallography supposed a fixed
structure, NMR structure investigations were made in solution.[29] Wüthrich eventually
received the 2002 Nobel Prize for Chemistry, “for his development of nuclear
magnetic resonance spectroscopy for determining the three-dimensional structure
of biological macromolecules in solution”.[30]
Editor-in-Chief of the Journal of Biomolecular NMR, Wüthrich is also best known for his
application of NMR to the study of prions. In 1996, he proposed a solution for
the tertiary structure of mouse PrP that conflicted
with the structure given by Prusiner at that time.[31] Wüthrich found that mouse
PrPc (121-131) “contains a two-stranded antiparallel beta-sheet and
three alpha-helices” (Riek et al. 1996: 180).[32] More than a third of his
short paper was devoted to graphic representations and the range of
representation modes was quite impressive:
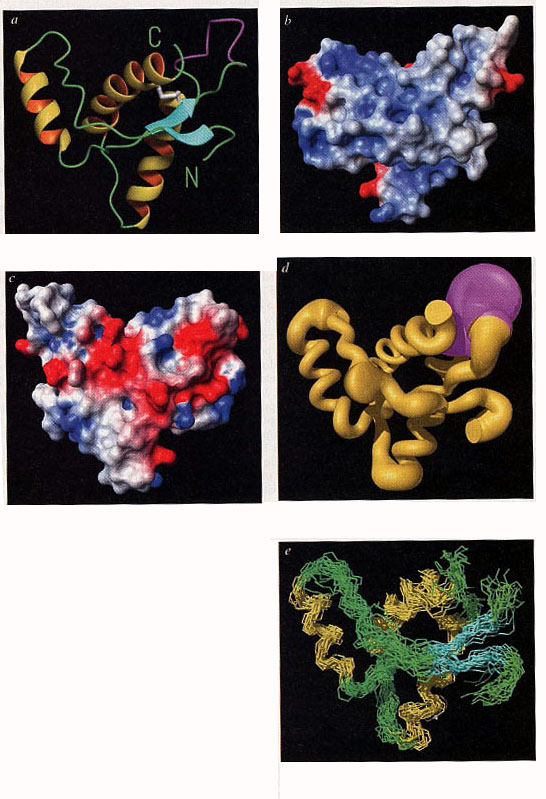
Figure 13 (from Riek et al. 1996: 181)
Crucially, a comparison with the mutation points identified in the
primary structure of human PrP supported the proposed
structure. From that time on NMR has been widely used in the study of prions
and we have seen that Prusiner has also used it.[33] In a short history
article published in 2001, Wüthrich confidently claimed: “we may soon be able to obtain information on the
structure of the disease-related, aggregated form of the prion protein” (Wüthrich
2001: 925).
An important aspect of the 1996 paper on the structure of the mouse prion
(Riek et
al. 1996) was the use of bioinformatics. In the box where ‘methods’
were described, we read that “the program Molmol
was used to generate the figure.” The authors referred to a previous
publication in the Journal of Molecular
Graphics that described Molmol
as a visualization device “with special emphasis on nuclear magnetic resonance
(NMR) solution structures of proteins” (Koradi et al. 1996: 51).[34] The classical approach of
biochemistry had thus been complemented by structural biology, with its emphasis
on 3D molecular structure.
In 1997, Wüthrich also used Molmol
to design a monoclonal antibody that could be used to establish diagnosis
(Corth et al. 1997). Three years
later, in 2000, Molmol helped to
visualize the human prion structure obtained by NMR (Zahn et al. 2000). A technical culture specific to computational biology
is embodied in all software designed to represent structures. Progressively,
this blurs the distinction between crystallographic and NMR methods: attention
is paid to the structure provided by
visualization devices irrespective of its mode of production. Moreover,
whereas crystallography was dominant in the static approach, researchers have
now started to use it for the identification of mobile parts of molecules.
Though scientists still belong to one or the other scientific culture, this
further blurs the difference between the two techniques.
Today, an image of the three-dimensional structure of human PrPc
stands on its own, with no caption, on Wüthrich’s homepages.[35] The knowledge of PrP
structure led to reification and just as the double helix stands for DNA, the
structure of PrPc now stands for the
prion.
IV-2. Salvation in the ‘yeast prion’?
.
In the same way as the bacterium Eschirechia
coli served as the typical prokaryote organism (organism without nucleus)
for the development of early molecular biology in the 1940s[36], the yeast Saccharomyces cerevisiae has now become
a useful organism for the development of a prion model. Yeast reproduces within
a few hours and is thus much easier to handle than mammalian prions. Scientists assume that the understanding of the
conformational change in yeast will provide valuable insight for studying
mammalian PrP.
Since the mid-1980s a journal called Yeast
has been devoted to these microorganisms. The Yeast editor for
In order to deepen the analysis of the three-dimensional structure,
Melki and co-workers tried to crystallize Ure2, which had been purified to
homogeneity. Since the characteristics of the beam line of the synchrotron they
had access to were incompatible with the size of their crystals, they used the
European Synchrotron Radiation Facility in
The fact that Melki and Wickner arrived independently at a similar
structure almost at the same time gives us an opportunity to see how the
visualization cultures attached to crystallography (Melki) and to NMR (Wickner)
have merged. Conventions have stabilized and biologists search in the same
databases from protein with similar structures. It is thus possible to compare
their respective representations of the Ure2p dimmer (Figure 14) and their
respective superimpositions of this protein with E. coli or A. thaliana
GST (Figure 15):
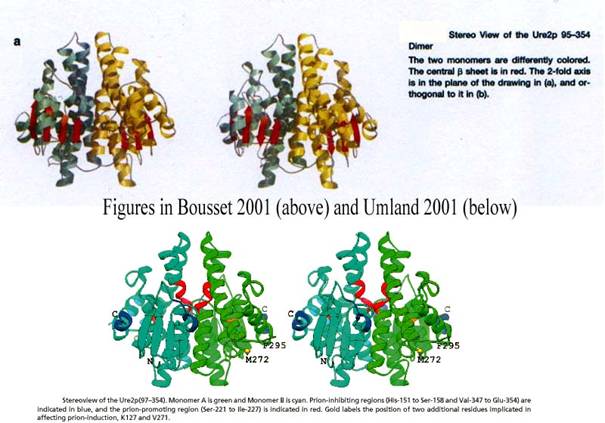
Figure 14 cyan
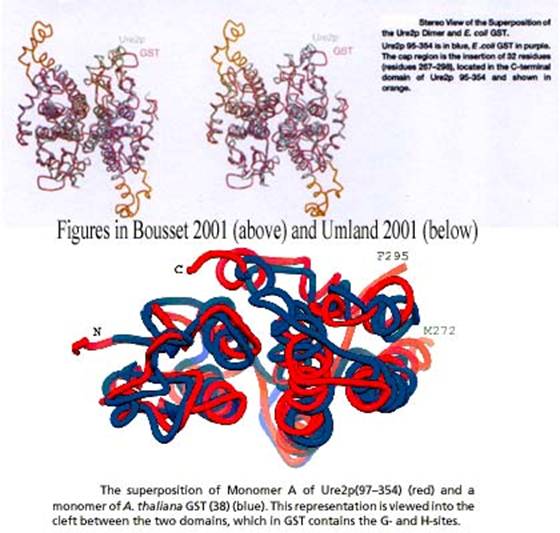
Figure 15
The use of large-scale technical systems such as a synchrotron became
embodied in representations and Melki’s structure featured on the cover of the
journal that published his paper (Bousset et
al. 2001). At this point three-dimensional structures of prions, here of
yeast, became emblematic of techniques, here of crystallography and the
synchrotron:
.

Figure 16 (Structure, 9,
This figure represented three dimers and the caption explained that this
“should help us to understand the mechanism of the amyloid formation associated
with a number of degenerative diseases.” The underlying motivation of the race
to the tertiary structure is to design drugs that can interfere with the
structure to avoid aggregation. In this sense, graphic representations not only
contributed to turning the prion hypothesis into a model, they also have a real
heuristic power that should soon be appreciated.
This being said, if research on [URE 3] and [PSI+] in Saccharomyces cerevisiae has become
paradigmatic of the prion model, some important considerations are sometimes
missing from the modelisation procedures. To begin with, no pathogenic effect
has ever been noticed in yeast. One of the two phenotypes that have been
studied presents interesting aggregation properties similar to that of PrPsc
but does not cause disease. If Ure2p can aggregate in vitro, it has now been proven that it does not cause [URE3] in vivo. Moreover, no homology sequence
has been found between mammalian and yeast prions. To encapsulate these
differences with mammalian prions, the term ‘propagon’ has recently been
proposed to designate yeast prions[45].
Conclusion
The present study has shown that in the early days the lack of
representation of the three-dimensional structure of the prion protein, which
is necessary to understand how it can convert into its pathogenic form,
restricted the credibility of the prion hypothesis. As the French TSE expert
Dominique Dormont put it:
“A scientific concept which is not supported by
direct visualization is always difficult to establish, whatever its origin may
be. In biology (I don’t speak about physics or mathematics), something you
cannot visualize always poses a lot of problems”.[46]
Yet, with the later work of Prusiner, Wüthrich, Wickner and Melki on
prion structures, representations progressively became the core of the prion
theory. In a recent paper on the “structure and assembly properties of the
yeast prion Ure2”, the word ‘picture’ has acquired a new meaning. The authors
write that they hope to get a “full picture of the molecular events at the origin
of prion propagation” (Bousset et al.
2002: 6). Thanks to computer graphics allied to NMR or, in this case,
crystallography, the epistemological function of visualization has moved from a
mere illustration to a possible explanation of the very nature of prions.
Visualization has played an important role in changing the status of the
prion concept from hypothesis to model. If biochemical experiments are still
paramount, computer graphics have been essential to determine the three-dimensional
structure of PrPc and are likely to play a
similar role in determining the structure of PrPsc. The development
of therapeutics will conclusively establish the importance of three-dimensional
structures since drug design aims at producing a molecule that can interfere
with the structure of the pathological protein.
More generally, now that the Human Genome Project has been completed,
there is little doubt that protein studies will continue to greatly benefit
from the development of the prion hypothesis and its visualization culture.
References
Aguzzi, A. et
al. (1997), “Neuro-immune connection in spread of
prions in the body?”, The Lancet 349:
742-743
Aigle, Michel & François Lacroute (1975), “Genetical aspects of [URE3], a non-mitochondrial, cytoplasmically inherited mutation in yeast”, Mol Gen Genet 136: 327-35
Amann, Klaus and Karin Knorr-Cetina
(1990), “The fixation of (visual) evidence”, in M. Lynch and
Anfinsen, Christian (1973), “Principles that
Govern the Folding of Protein Chains”, Science
181: 223-30
Appel, R.D. et
al. (1994), “A new generation of information retrieval tools for
biologists: the example of the ExPASy WWW server” Trends Biochem. Sci. 19: 258-260
Baldwin, M.A. et al. (1998), “The three-dimensional structure of prion protein:
implications for prion disease”, Biochemical
Society Transactions 26: 481-486.
Bousset, Luc et al. (2001), “Structure of the globular region of the prion
protein Ure2 from the yeast Saccharomyces cerevisiae”, Structure (Camb) 9(1): 39-46
Bousset, Luc et al. (2002), “Structure and assembly properties of the yeast
prion Ure2”, C.R. Biologies
325: 3-8
Brandner, S. et al. (1996), “
Cambrosio, Alberto (2000), “Argumentation, représentation, intervention: les rôles
de l’imagerie dans les discours scientifiques”, ASp 27/30: 95-112
Chamak, Brigitte “Prion Research and the
Public Sphere in France”, this volume
Chernoff, Y.O. et al. (1995), “Role of the chaperone protein Hsp104 in propagation
of the yeast prion-like factor [psi+]”, Science 268(5212): 880-884.
Corth, C. et
al. (1997), “Prion (PrPSc)-specific epitope
defined by a monoclonal antibody”, Nature
390(6655), 74-77
Cox, B.S (1965), “PSI, a cytoplasmic
suppressor of super-suppressor in yeast”, Heredity
20: 505-521
Couzin, Jennifer (2002), “In Yeast,
Prions’ Killer Image Doesn’t Apply”, Science
297: 758-761
Crick, Francis (1958), “On Protein Synthesis”,
in F.K. Sanders (Ed.), The Biological
Replication of Macromolecules,
Donne, D.G. et
al (1997), “Structure of the recombinant full-length hamster prion protein
PrP(29-231): the N terminus is highly flexible”, PNAS USA 94(25): 13452-7
Dressel, Kerstin, “Paradigm Change? Explaining
the Nature of the TSE Agent in Germany” , this volume
Fernandez-Bellot, E.
& C. Cullin (2001), “The protein-only theory and
the yeast saccharomyces cerevisiae: the prions and the propagons.” Cell Mol
Life Sci. 58(12-13): 1857-78
Ferrin, T.E. et al. (1988), “The MIDAS Display System”, J. Mol.
Graphics 6(1): 13-27, 36-37
Francoeur,
Eric (2001), “Molecular Models and the Articulation of Structural Constraints
in Chemistry”, in U. Klein (Ed.) Tools and Modes of Representation in the
Laboratory Sciences,
Francoeur, Eric & Segal, Jérôme (2004), “From model kits to interactive computer graphics”,
in S. de Chadarevian and
Gasset, M. et
al. (1992), “Predicted alpha-helical regions of the prion protein when
synthesized as peptides form amyloid”, PNAS USA 89(22), 10940-4
Glatzel, M. and A. Aguzzi (2000), “PrPc
expression in the peripheral nervous system is a determinant of prion
neuroinvasion”, Journal of General
Virology 81: 2813–2821
Glockshuber, R. et al. (1997), “Three-dimensional NMR structure of a self-folding
domain of the prion protein PrP(121-231), Trends
in Biochemical Sciences 22(7): 241-242
Griesemer, James
R. (1991), "Must Scientific Diagrams Be Eliminable? The Case of Path
Analysis," Biology and Philosophy
6: 155-180
Güntert, P. et al. (1991), “Efficient computation of three-dimensional
protein structures in solution from nuclear magnetic resonance data using the
program DIANA and the supporting programs CALIBA, HABAS and GLOMSA”, J. Mol.
Biol. 217: 517-30
Huang, Z. et
al. (1994), “Proposed three-dimensional structure for the cellular prion
protein”, PNAS USA 91(15): 7139-43
Huang, Z. et al. (1996), “Structures of prion proteins and conformational
models for prion diseases”, Curr Top
Microbiol Immunol. 207: 49-67
James, T.L. et
al. (1997), “Solution structure of a 142-residue recombinant prion protein
corresponding to the infectious fragment of the scrapie isoform”,
PNAS USA 94(19): 10086-91
Johnson Carroll K (1965), “OR TEP: A FORTRAN
Thermal-Ellipsoid Plot Program for Crystal Structure Illustrations”, ONRL Report #3794. Oak Ridge, Ten., Oak
Ridge National Laboratory
Kaneko K. et
al. (1997), “Evidence for protein X binding to a discontinuous epitope on the cellular prion protein during scrapie prion
propagation”, PNAS USA 94: 10069-10074
Kay, Lily E. (2000), Who Wrote the Book of Life? A History of the Genetic Code, Chicago:
University of
Keyes, Martha E. (1999), “The Prion Challenge
to the ‘Central Dogma’ of Molecular Biology, 1965-1991. Part I: Prelude to
Prions” & “Part II: The Problem with Prions”, Studies in History and Philosophy of Biological and Biomedical Sciences,
30 (1&2): 1-19 & 181-218
Kim, Ki-Heung, “Styles
of Scientific Practice and the Prion Controversy”, this volume
Koradi, R. et al. (1996), “MOLMOL: a program for display and analysis of
macromolecular structures”, Journal of
Molecular Graphics 14(1): 51-55
Kuhn, Thomas
S. (1977), The Essential Tension:
Selected Studies in Scientific Tradition and Change. Chicago: University of
Chicago Press
Lacroute, François (1971), “Non-Mendelian mutation allowing ureidosuccinic
acid uptake in yeast”, J Bacteriol 106: 519-22
Lakoff, George and Mark Johnson (1980), Metaphors We Live By. Chicago:
Latour, Bruno and Steve Woolgar
(1979), Laboratory Life: The Social
Construction of Scientific Facts. Beverly Hills: Sage Publications
Latour,
Bruno
(1993), "Le "Pédofil" De Boa Vista -
Montage Photo-Philosophique," in La Clef De Berlin. Paris: La Découverte: 171-225
Levinthal, Cyrus (1966), “Molecular
Model-Building by Computers”, Scientific
American 214: 42-52
Lynch, Michael
(1985), "Discipline and the Material Form of Images," Social Studies of Science 15 (1): 37-66
Lynch, Michael
(1998), "The Production of Scientific Images: Vision and Re-Vision in the
History, Philosophy, and Sociology of Science," Communication and Cognition 31, no. 2/3. 213-28
Martz, Eric and Francoeur,
Eric (2001) “History of Visualization of Biological Macromolecules”, http://www.umass.edu/microbio/rasmol/history.htm
(latest revision 9/2001)
Maunoury, M.T. et al. (1999), “Observer la science en action ou, comment
les sciences de l’information permettent de suivre l’évolution et la
convergence des concepts de prion et d’hérédité non mendélienne dans la
littérature”, Médecine/Sciences 15(4):
577-82
Merz, Patricia, et
al. (1981) “Abnormal Fibrils from Scrapie-infected Brain”, Acta Neuropathologica 54: 63-74
Merz, Patricia et
al. (1984) “Infection-Specific Particle from the Unconventional Slow Virus
Diseases”, Science 225: 437-440.
Morange, Michel (1998), A History of Molecular Biology,
Ottiger, M. et al. (1994) “The NMR solution
structure of the pheromone Er-2 from the ciliated protozoan Euplotes
raikovi”, Protein
Science 3: 1515-1526
Pauling, Linus et al. (1951), “The Structure of Proteins: Two Hydrogen-Bonded
Helical Configurations of the Polypeptide Chain”, PNAS
Poulsen, Maj-Britt
Juhl & Hanne Andersen, “The
Early History of the Protein-Only Hypothesis
Scientific Change and Multidisciplinary
Research”, this volume
Prusiner,
Stanley (1982), “Novel Proteinaceous
Infectious Agent Particles
Cause Scrapie”, Science 216: 136-144
Prusiner, Stanley et al. (1983), “Scrapie prions aggregate to form amyloid-like birefringent rods”, Cell
35: 349-358
Prusiner,
Prusiner, Stanley et al. (1987), “On the biology of prions”, Acta Neuropathologica 72: 299-314
Prusiner,
Prusiner,
Prusiner,
Prusiner, Stanley (1998), “Prions”, PNAS USA 95(23): 13363-83
Prusiner, Stanley et al. (1987), “On the biology of prions”, Acta Neuropathol (Berl)
72(4): 299-314
Raeber, A.J. et al. (1998), “Transgenic and Knockout Mice in Research on Prion
Diseases”, Brain Pathology 8: 715-733
Rheinberger, Hans-Jörg
(1997), Toward a History of Epistemic
Things: Synthesizing Proteins in the Test Tube (Writing Science). Stanford:
Stanford
Riek, R. et
al. (1996), “NMR structure of the mouse prion protein PrP(121-231)”, Nature 382: 180-182
Riek, R. et
al. (1997), “NMR characterization of the full-lenght
recombinant murine prion protein mPrP
(23-231)”, FEBS letters 413: 282-288
Rudwick, Martin
J. S. (1976), "The Emergence of a Visual Language for Geological
Science," History of Science 14:
149-95
Sarkar, Sahotra
S. (1996), “Thinking of Biology - Decoding ‘coding’ - information and DNA”, Bioscience 46: 857-64
Segal, Jérôme (2002), “Les premiers ‘replieurs’ français: Michel
Goldberg à l’Institut
Pasteur et Jeannine Yon à Orsay”,
Revue pour l’histoire
du CNRS
Shulamn, Robert G. (2000), “D.C. Phillips”,
Biographical Memoirs of the National
Academy of Sciences (National Academy Press, 2000, 166-181 or http://books.nap.edu/books/030907035X/html/166.html)
Soojung-Kim Pang, Alex (1997), "Visual Representation and Post-Contructivist History of Science," Historical Studies in the Physical and
Biological Sciences 28: 139-71.
Thieffry, Denis (1996), “E. coli as a
model system with which to study cell-differentiation”, Hist. Philos. Life Sci.
18: 163-193
Thual, Carine et al. (1999), “Structural
characterization of Saccharomyces cerevisiae prion-like protein Ure2”, J Biol Chem 274(19):13666-74
Umland, T.C. et al. (2001), “The crystal structure of the nitrogen regulation
fragment of the yeast prion protein Ure2p”, PNAS
USA 98(4): 1459–1464
Wickner, Reed B. (1994), “[URE3] as an altered
URE2 protein: evidence for a prion analog in Saccharomyces cerevisiae”, Science 264(5158): 566-9
Wickner, Reed B. et al. (1995), “[PSI] and [URE3] as yeast prions”, Yeast 11(16): 1671-85
Wüthrich, Kurt (2001), “The way to NMR
structures of proteins”, Nature
Structural Biology 8: 923-925
Zahn, R. et
al. (2000), “NMR solution structure of the human prion protein”, PNAS USA 97(1): 145-50
